DeepGBoost
Machine learning algorithm based on gradient boosting forest that merges the power of tree ensembles with neural network architectures.
Algorithm
DeepGBoost implements the Distributed Gradient Boosting Forest (DGBF), introduced in:
Delgado-Panadero, A., Benitez-Andrades, J. A., & Garcia-Ordas, M. T. (2023). A generalized decision tree ensemble based on the NeuralNetworks architecture: Distributed Gradient Boosting Forest (DGBF). Applied Intelligence, 53, 22991-23003. https://doi.org/10.1007/s10489-023-04735-w
Classical tree ensemble methods — RandomForest (bagging) and GradientBoosting (boosting) — cannot perform hierarchical representation learning the way neural networks do. DGBF addresses this by combining both approaches into a unified formulation that defines a graph-structured tree ensemble with distributed representation learning, without requiring back-propagation or parametric models.
The ensemble prediction is:
$$F(x) = \sum_{l=1}^{L} RF_l(x) = \frac{1}{T} \sum_{l=0}^{L} \sum_{t=0}^{T} h_{l,t}(x)$$
where L is the number of boosting layers and T is the number of trees per layer. Each RandomForest layer is the analogue of a dense network layer, with distributed gradients replacing back-propagation.
RandomForest (L = 1) and GradientBoosting (T = 1) are recovered as special cases.
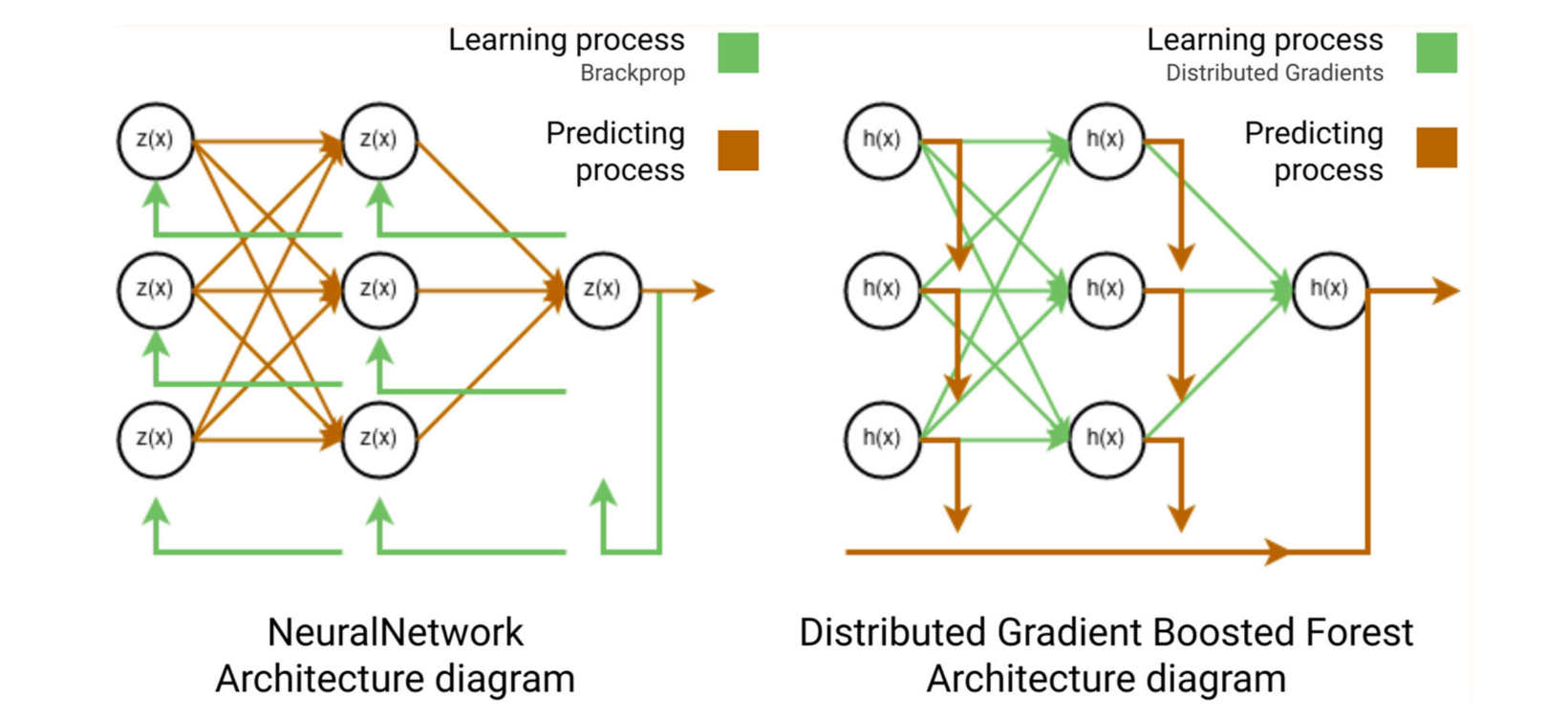
Fig. 1 — NeuralNetwork vs DGBF architecture
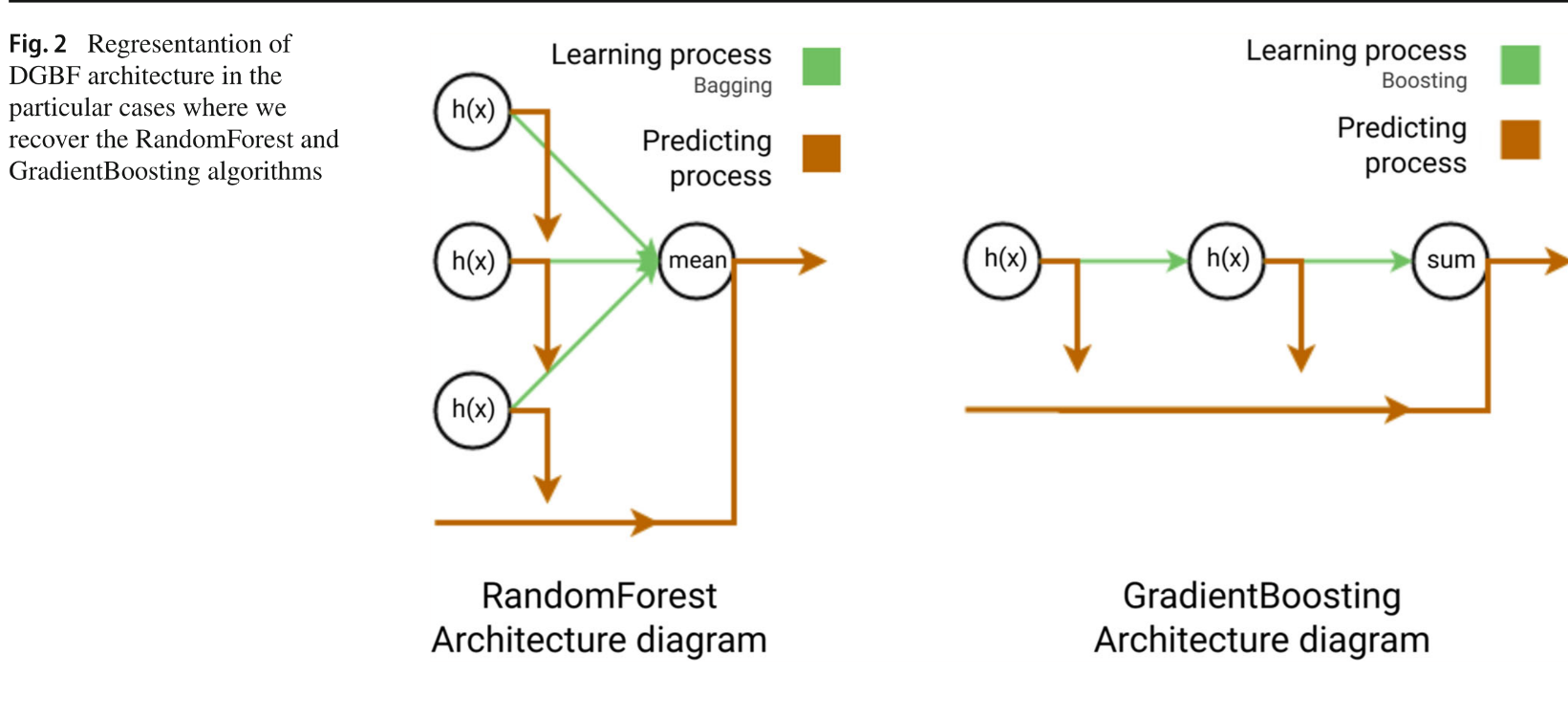
Fig. 2 — RandomForest & GradientBoosting as DGBF special cases
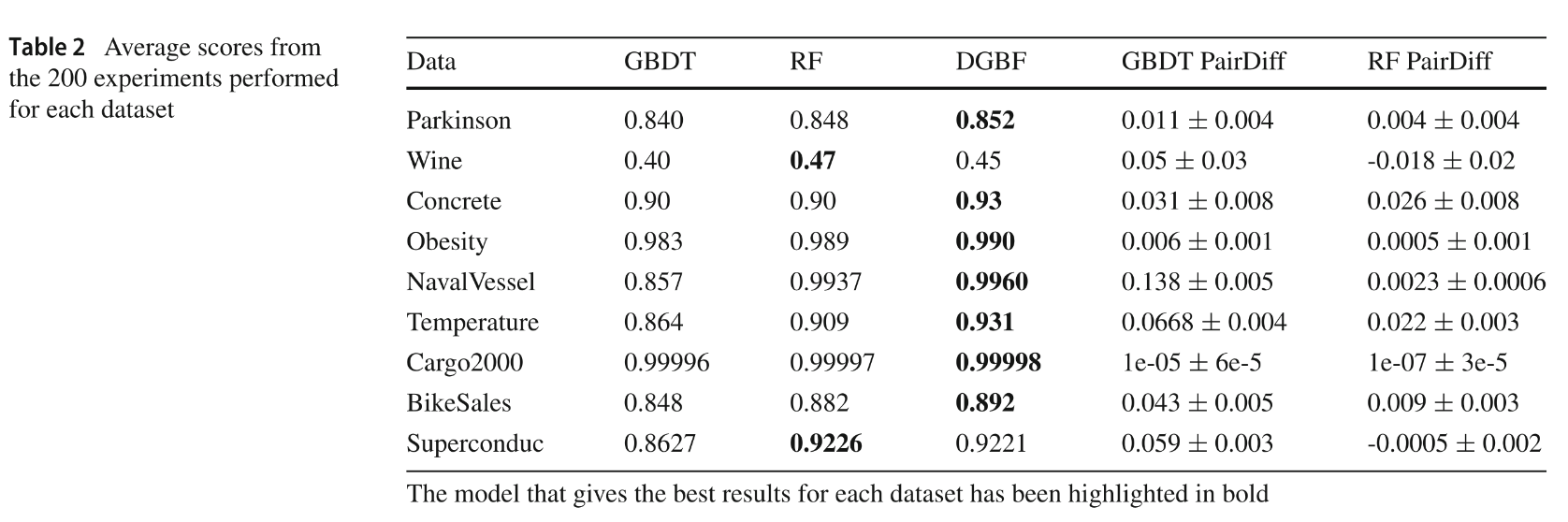
Fig. 3 — Benchmark results across 9 UCI datasets
Installation
pip install deepgboost
Optional plotting support:
pip install deepgboost[plotting]
Install from source with development dependencies:
git clone https://github.com/delgadopanadero/deepgboost.git
cd deepgboost
pip install -e ".[dev]"
Quick Start
Regression
from sklearn.datasets import load_diabetes
from sklearn.model_selection import train_test_split
from deepgboost import DeepGBoostRegressor
X, y = load_diabetes(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
model = DeepGBoostRegressor(n_trees=10, n_layers=10, max_depth=4, learning_rate=0.1)
model.fit(X_train, y_train)
print(model.score(X_test, y_test))
Classification
from sklearn.datasets import load_breast_cancer
from deepgboost import DeepGBoostClassifier
X, y = load_breast_cancer(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
clf = DeepGBoostClassifier(n_trees=10, n_layers=10, learning_rate=0.1)
clf.fit(X_train, y_train)
print(clf.score(X_test, y_test))
Early stopping with callbacks
from deepgboost import DeepGBoostRegressor, EarlyStoppingCallback, EvaluationMonitorCallback
model = DeepGBoostRegressor(n_trees=5, n_layers=20, learning_rate=0.05)
model.fit(
X_train, y_train,
eval_set=[(X_test, y_test)],
callbacks=[EvaluationMonitorCallback(period=5), EarlyStoppingCallback(rounds=10)],
)
Citation
@article{delgado2023dgbf,
author = {Delgado-Panadero, {\'A}ngel and Ben{\'i}tez-Andrades, Jos{\'e} Alberto and Garc{\'i}a-Ord{\'a}s, Mar{\'i}a Teresa},
title = {A generalized decision tree ensemble based on the {NeuralNetworks} architecture: {Distributed Gradient Boosting Forest (DGBF)}},
journal = {Applied Intelligence},
volume = {53},
pages = {22991--23003},
year = {2023},
doi = {10.1007/s10489-023-04735-w}
}
